Note
Click here to download the full example code
Neuropythy and Neuralink: Patient specific visual field maps based on MRI¶
Neuropythy [Benson2018] is a python package that predicts patient-specific visuotopies based on MRI scans of the human visual cortex. It is possible to use Neuropythy within pulse2percept as the visual field map for cortical models.
First, make sure neuropythy is installed, which can be done with
pip install neuropythy.
Creating a Neuropythy visual field map¶
NeuropythyMap requires a subject parameter,
which can be either:
1) ‘fsaverage’ (the average of freesurfer subjects)
2) a string with a subject from the Benson Winawer 2018 dataset (‘S1201’-‘S1208’)
3) the path to a freesurfer subject directory in a format neuropythy can load
4) the name of a subject in any directory specified by ny.config['freesurfer_subject_paths'] or
5) an instantiated neuropythy.mri.core.Subject
If you do not have it downloaded already, the Benson Winawer 2018 dataset will
be automatically downloaded and cached to the provided cache_dir parameter,
which defaults to ~/.neuropythy_p2p. If you have already downloaded the dataset,
elsewhere, just add the subjects folder of the dataset to ny.config['freesurfer_subject_paths'] and p2p
will be able to load subjects from your predownloaded directory.
import pulse2percept as p2p
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings("ignore") # ignore matplotlib warnings
# this could take a while if you haven't already downloaded the dataset
nmap = p2p.topography.NeuropythyMap(subject='fsaverage', regions=['v1'])
nmap
NeuropythyMap(cort_nn_thresh=1000, jitter_boundary=False,
jitter_thresh=0.3, left_offset=None, ndim=3,
regions=['v1'])
NeuropythyMap provides a number of methods to transform visual field coordinates into cortical coordinates:
In contrast to other visual field maps, you may also specify a cortical surface that the visual field coordinates should be mapped to. By default, the cortical surface is set to ‘midgray’, but you can also specify ‘white’ or ‘pial’, or other surfaces accepted by neuropythy.
Lets use our map in a model and visualize the transform
model = p2p.models.cortex.ScoreboardModel(vfmap=nmap, regions=['v1'])
model.build()
fig, axes = plt.subplots(ncols=2, figsize=(10, 4))
model.plot(style='cell', ax=axes[0])
model.plot(style='scatter', ax=axes[1])
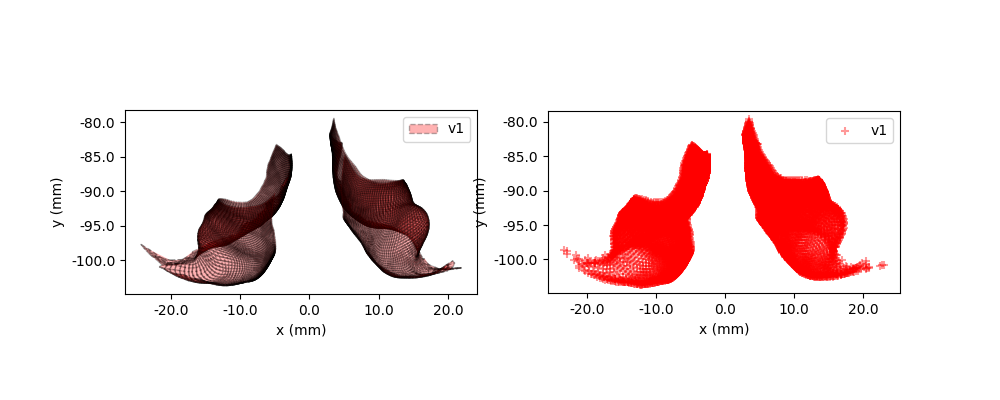
Warning: Plotting 2D projection of 3D data. You might want plot3D() instead
Warning: Plotting 2D projection of 3D data. You might want plot3D() instead
<Axes: xlabel='x (mm)', ylabel='y (mm)'>
Note, if we call the usual plot() method, it will be plotting a 2D projection of the points. This can look quite strange sometimes, since the points are inherently 3D. To plot the points in 3D, we can use the plot3D() method.
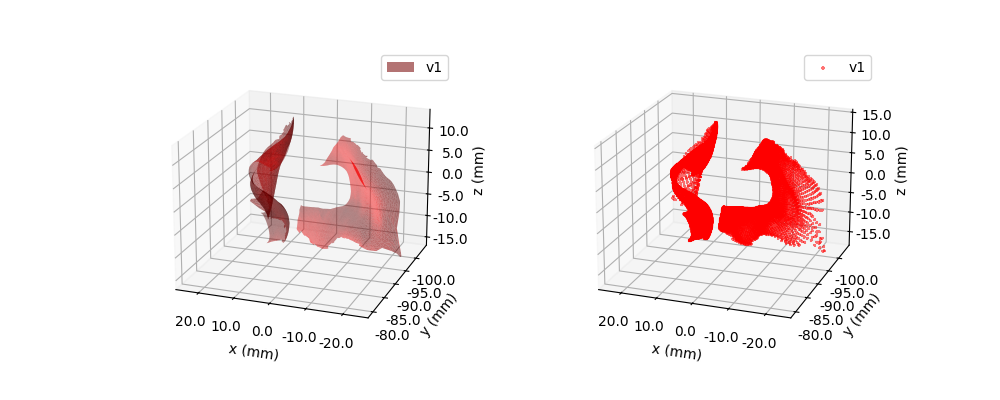
<Axes3D: xlabel='x (mm)', ylabel='y (mm)', zlabel='z (mm)'>
Neuropythy can use up to ‘v1’, ‘v2’, ‘v3’. If you want to use all three regions, you can specify them as a list.
Lets use all three regions and plot the result (note it can get a little messy):
nmap = p2p.topography.NeuropythyMap(subject='fsaverage', regions=['v1', 'v2', 'v3'])
model = p2p.models.cortex.ScoreboardModel(xrange=(-20, 20), yrange=(-20, 20), xystep=0.5,
vfmap=nmap, regions=['v1', 'v2', 'v3'])
fig = plt.figure(figsize=(10, 4))
ax1 = fig.add_subplot(121, projection='3d')
model.plot3D(ax=ax1, style='cell')
ax2 = fig.add_subplot(122, projection='3d')
model.plot3D(ax=ax2, style='scatter')
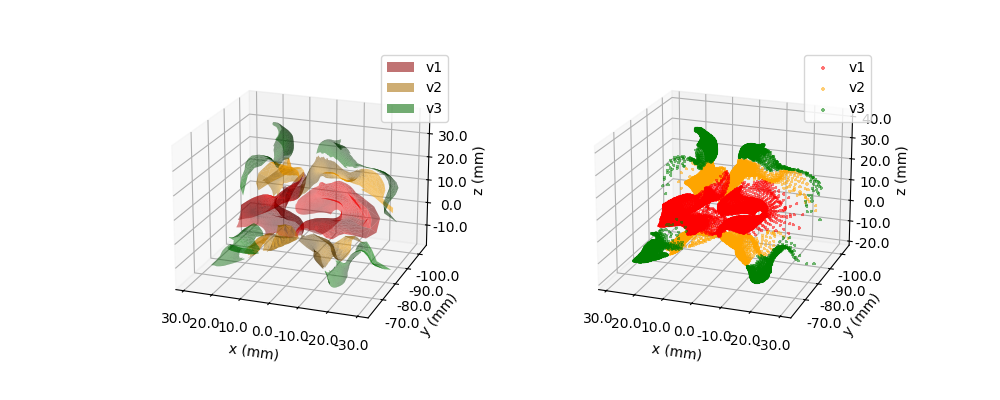
<Axes3D: xlabel='x (mm)', ylabel='y (mm)', zlabel='z (mm)'>
Note that the regions are not necessarily
contiguous, and the map will likely be discontinuous at the boundaries between
visual areas. To combat this, you can set the jitter_boundary to True
to add a small amount of noise to the boundary to make it less noticeable.
This is turned off by default.
Placing implants with neuropythy maps¶
When using a cortical map, it is important to place your implant in the correct 3D location; the default z value of 0 will likely not be on a cortical surface.
For the Neuralink implant, there exists a helper function,
from_neuropythy, which will
automatically place neuralink threads at specified visual field locations across
the cortical surface. Threads will be inserted perpendicular to the cortical surface
up to rand_insertion_angle. Similar helper functions are under development
for other cortical implants.
Lets place a Neuralink implant across the right hemisphere of the cortex:
nmap = p2p.topography.NeuropythyMap(subject='fsaverage', regions=['v1'])
model = p2p.models.cortex.ScoreboardModel(vfmap=nmap, xrange=(-4, 0), yrange=(-4, 4), xystep=.25).build()
nlink = p2p.implants.cortex.Neuralink.from_neuropythy(nmap, xrange=model.xrange, yrange=model.yrange,
xystep=1,rand_insertion_angle=0)
print(len(nlink.implants), " total threads")
fig = plt.figure(figsize=(10, 4))
ax1 = fig.add_subplot(121, projection='3d')
model.plot3D(ax=ax1, style='cell')
nlink.plot3D(ax=ax1)
ax2 = fig.add_subplot(122)
model.plot(style='cell', ax=ax2)
nlink.plot(ax=ax2)
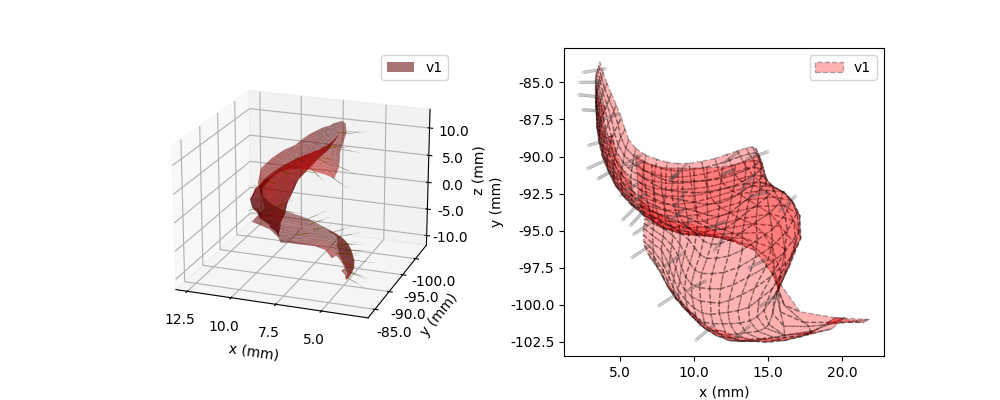
36 total threads
Warning: Plotting 2D projection of 3D data. You might want plot3D() instead
<Axes: xlabel='x (mm)', ylabel='y (mm)'>
Finally, lets predict what a percept would look like if we stimulated one electrode on each thread, using the scoreboard model:
nmap = p2p.topography.NeuropythyMap(subject='fsaverage', regions=['v1'], jitter_boundary=True)
model = p2p.models.cortex.ScoreboardModel(rho=800, xrange=(-15, 15), yrange=(-15, 15), xystep=.25, vfmap=nmap).build()
nlink = p2p.implants.cortex.Neuralink.from_neuropythy(nmap, xrange=model.xrange, yrange=model.yrange,
xystep=3,rand_insertion_angle=0)
nlink.stim = {e : 1 for e in [i for i in nlink.electrode_names if i[-1] == '1']}
percept = model.predict_percept(nlink)
percept.plot()
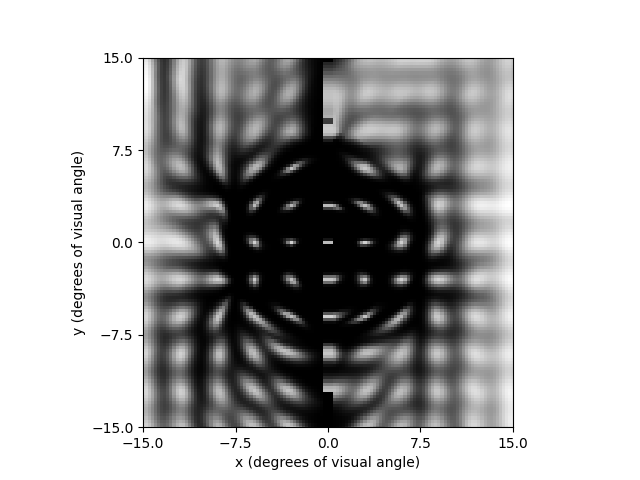
<Axes: xlabel='x (degrees of visual angle)', ylabel='y (degrees of visual angle)'>
Total running time of the script: ( 3 minutes 13.347 seconds)