Computational Models¶
The models module provides a number of published
and verified computational models that can be used to predict neural responses
or visual percepts resulting from electrical stimulation.
A Model object consists of:
- a
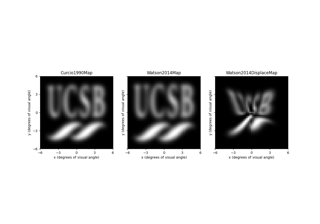
SpatialModel, describing how electrical stimulation affects the neural tissue or elicited phosphene in different spatial locations of the visual field, and/or - a
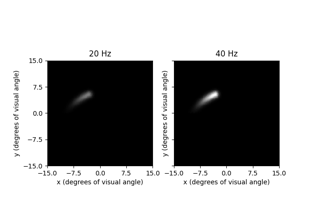
TemporalModel, describing how the response of the neural tissue or elicited phosphene evolves over time.
pulse2percept provides the following computational models:
| Reference | Model | Type |
| generic | FadingTemporal |
temporal |
| [Thompson2003] | Thompson2003Model |
spatial |
| [Thompson2003] | Thompson2003Spatial |
spatial |
| [Horsager2009] | Horsager2009Model |
temporal |
| [Horsager2009] | Horsager2009Temporal |
temporal |
| [Nanduri2012] | Nanduri2012Model |
spatial + temporal |
| [Nanduri2012] | Nanduri2012Spatial |
spatial |
| [Nanduri2012] | Nanduri2012Temporal |
temporal |
| [Beyeler2019] | AxonMapModel |
spatial |
| [Beyeler2019] | ScoreboardModel |
spatial |
| [Granley2021] | BiphasicAxonMapModel |
spatial + temporal |
Note
Spatial and temporal models can be mix-and-matched to create new models. See Creating your own model.
Basic usage¶
All models follow the same basic work flow:
- Initialize the model with the desired model parameters.
- Build the model to perform one-time heavy computations such as building
the axon map in
AxonMapModel. - Predict a percept by passing an implant that contains a stimulus. The
model will return a
Perceptobject that acts as a data container with labeled axes.
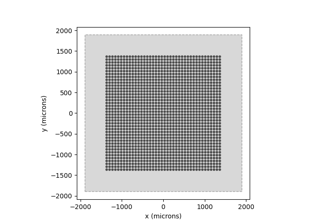
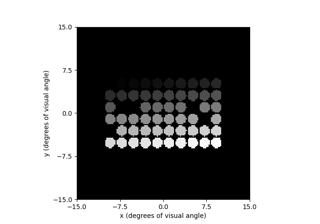
Here is how to run the ScoreboardModel:
# Initialize the model:
In [1]: from pulse2percept.models import ScoreboardModel
In [2]: model = ScoreboardModel(rho=200)
# Build the model:
In [3]: model.build()
Out[3]:
ScoreboardModel(engine=None, grid_type='rectangular',
n_gray=None, n_jobs=1, n_threads=2,
noise=None, retinotopy=Watson2014Map,
rho=200, scheduler='threading',
spatial=ScoreboardSpatial, temporal=None,
thresh_percept=0, verbose=True,
xrange=(-15, 15), xystep=0.25,
yrange=(-15, 15))
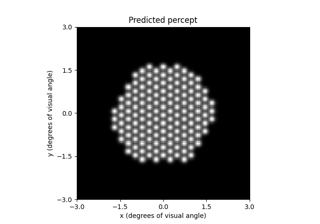
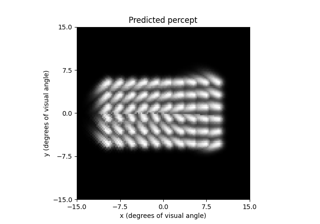
# Predict the percept resulting from stimulating Electrode
# A8 in Argus II with 30 uA:
In [4]: from pulse2percept.implants import ArgusII
In [5]: percept = model.predict_percept(ArgusII(stim={'A8': 30}))
Building your own model¶
To build your own model, you can mix and match spatial and temporal models at will.
For example, to create a model that combines the scoreboard model described in [Beyeler2019] with the temporal model cascade described in [Nanduri2012], use the following:
# Instantiate:
model = Model(spatial=ScoreboardSpatial(),
temporal=Nanduri2012Temporal())
# Build:
model.build()
# etc.
To create a more advanced model, you will need to subclass the appropriate base
class. For example, to create a new spatial model, you will need to subclass
SpatialModel and provide implementations for
the following methods:
dva2ret: a means to convert from degrees of visual angle (dva) to retinal coordinates (microns).ret2dva: a means to convert from retinal coordinates to dva._predict_spatial: a method that accepts anElectrodeArrayas well as aStimulusand computes the brightness at all spatial coordinates ofself.grid, returned as a 2D NumPy array (space x time).
In addition, you can customize the following methods:
__init__: the constructor can be used to define additional parameters (note that you cannot add parameters on-the-fly)get_default_params: all settable model parameters must be listed by this method_build(optional): a way to add one-time computations to the build process
A full working example:
class MySpatialModel(SpatialModel):
def __init__(self, **params):
"""Constructor"""
# Make sure to call the parent's (SpatialModel's constructor):
super(MySpatialModel, self).__init__(self, **params)
# You can set additional parameters here (e.g., stuff you will
# need later on in ``_build``). You will not be able to add
# parameters outside the constructor or ``get_default_params``.
self.n_fib = 100
def get_default_params(self):
"""Return a dictionary of settable model parameters"""
# Get all parameters already set by the parent (SpatialModel):
params = super(MySpatialModel, self).get_default_params()
# Add our own:
params.update(myparam=1)
# Return the combined dictionary:
return params
def dva2ret(self, dva):
"""Convert degrees of visual angle (dva) into retinal coords (um)"""
return 280.0 * dva
def ret2dva(self, ret):
"""Convert retinal corods (um) to degrees of visual angle (dva)"""
return ret / 280.0
def _build(self):
"""Perform heavy computations during the build process"""
# Perform some expensive computation using parameters you
# initialized in the constructor:
self.heavy = some_heavy_comp(self.n_fib)
def _predict_spatial(self, earray, stim):
"""Calculate the spatial response at different time points"""
resp = np.zeros(self.grid.size, stim.time.size)
for idx_t, t in enumerate(stim.time):
for idx_xy, (x, y) in enumerate(self.grid):
# Response at (x,y,t) is the sum of x,y coordinates and
# all the stimuli at time t (an arbitrary, silly choice):
resp[idx_xy, idx_t] = x + y + np.sum(stim[:, t])
return resp
Similarly, a new temporal model needs to subclass from
TemporalModel and provide a
_predict_temporal method:
class MyTemporalModel(TemporalModel):
def _predict_temporal(self, stim, t_percept):
"""Calculates the temporal response at different time points"""
# Response at (x,y,t) is the stimulus at (x,y,t). Use stim's smart
# indexing to do automatic interpolation:
return stim[:, t_percept]
Stand-alone models vs. spatial/temporal model components¶
In general, you will want to work with Model
objects, which provide all the necessary glue between a spatial and/or a
temporal model component. Objects are named accordingly:
- An object named *Model is based on
Model - An object named *Spatial is based on
SpatialModel - An object named *Temporal is based on
TemporalModel
However, nobody stops you from instantiating a spatial or temporal model directly:
# Option 1 (preferred): Work with Model objects:
from pulse2percept.models import Model, Nanduri2012Temporal
model = Model(temporal=Nanduri2012Temporal())
model.build()
model.predict_percept(implant)
# Option 2: Work directly with a temporal model:
model = Nanduri2012Temporal()
model.build()
model.predict_percept(implant.stim)
The differences between the two are subtle:
- As you can see from the example above, a temporal model will expect a
Stimulusobject in itspredict_perceptmethod (because it has no notion of space). It will return a 2-D NumPy array (space x time). - In contrast, the stand-alone model will expect a
ProsthesisSystemobject (which provides a notion of space and itself contains aStimulus), and will return aPerceptobject.
Getting and setting parameters¶
A Model will hide the complexity that some
parameters exist only in the spatial or temporal model component.
Consider the following model:
In [6]: from pulse2percept.models import (Model, ScoreboardSpatial,
...: Nanduri2012Temporal)
...:
In [7]: model = Model(spatial=ScoreboardSpatial(),
...: temporal=Nanduri2012Temporal())
...:
# Set `rho` param of the scoreboard model (works even though it's really
# `model.spatial.rho`):
In [8]: model.rho = 123
# Print the simulation time step of the Nanduri model (works even though
# it's really `model.temporal.dt`):
In [9]: print(model.dt)
0.005
Although rho exists only in the scoreboard model, and dt exists only
in the temporal model, you can get and set them as if they were part of the
main model.
Warning
If a parameter exists in both spatial and temporal models (e.g.,
thresh_percept), then calling model.thresh_percept = 0 will update
both the spatial and temporal model.
Alternatively, use model.spatial.thresh_percept = 0 or
model.temporal.thresh_percept = 0.